Union Minister Nitin Gadkari has grabbed headlines with his controversial assertion that Covid-19 was caused not by a natural virus but one that was manufactured in a Chinese laboratory. Gadkari’s claim marked the first time that an Indian government minister had remarked on the origin of the novel coronavirus. It’s not known whether this is his personal view or the official position of the Indian government. It should be noted, though, that Gadkari is not alone in making such claims.
US President Donald Trump has also asserted that this virus originated in a Chinese lab rather than the Huanan Seafood Wholesale Market in Wuhan, which Chinese authorities have pointed to as the likely origin of the virus. French Nobel Laureate Luc Montagnier, co-discoverer of the human immunodeficiency virus (HIV), has made similar assertions that the virus emerged from a lab in Wuhan. In fact, the world’s been abuzz with theories about the origin of the virus. But most scientists who have sifted through the evidence say it’s extremely unlikely that the virus came out of a laboratory.
Scientific evidence
Let’s go through the scientific evidence about the origins of the virus. In early January, China released the first genome sequence of a new virus, later called severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) or Covid-19. Since then over 25,000 SARS-CoV-2 genomic sequences have been fed into the Global Initiative on Sharing All Influenza Data (GISAID) database. These are being quickly made publicly available to help researchers understand the origin of this virus and to track its march across the world.
SARS-CoV-2 is the seventh coronavirus to infect humans, two of which are endemic and account for 20 per cent to 30 per cent of common colds annually. Prior to SARS-CoV-2, the most recent family members – SARS-CoV in 2002-03 and the Middle East Respiratory Syndrome coronavirus (MERS-CoV) in 2012, all of which cause acute respiratory distress -- originated in bats and jumped into humans via other animals – civet cats and camels, respectively. The closest genetic relatives of SARS-CoV-2 are coronaviruses that were isolated from bats (RaTG13) and pangolins, which are scaly skinned mammals.
SARS-CoV-2 enters human cells by first binding to a protein called ACE2 (angiotensin converting enzyme 2) present on the surface of these cells. Importantly, five key amino acid residues in the viral spike proteins that allow it to bind the human ACE2 receptor are consistent between SARS-CoV-2 and Pangolin-CoV, but not RaTG13. A key cleavage site within the spike protein, which is considered important for efficient infection and transmission, is found only in SARS-CoV-2 but not in either RaTG13 nor Pangolin-CoV. This supports a scenario in which SARS-CoV2, just like its predecessors, jumped into humans via an intermediate animal host, in this case pangolins, and continued to change in humans until it broke loose. This gradual change is indicative of the virus evolving in different species – bats, pangolins and humans.
In a non-peer reviewed study published on May 13 on aRxiv, a pre-print server, researchers from Australia combined in silico structural modelling and docking studies with available genomic and structural biology data to test how efficiently the SARS-CoV-2 spike protein binds to various animal ACE2 proteins. Their results suggest that SARS-CoV-2 is uniquely evolved to bind and infect human cells.
These studies also predict that SARS-CoV-2 will be inefficient in infecting other animals – hamsters, ferrets, felines, dogs and cattle _ _ a finding that should give some relief to pet owners and lovers of other animals. Occasional infections in some of these animals, such as cats and dogs, has already been documented.
Based on genomic features and structure-function studies, the SARS-CoV-2 spike protein appears to be optimised for binding to ACE2 on human cells. This would not have been predicted by computational analysis of the receptor-binding domain (RBD) in the spike protein from SARS-CoV. As a result, researchers conclude that the high-affinity binding of the SARS-CoV-2 spike protein to human ACE2 is most likely the result of natural selection involving a human or human-like ACE2. The presence of highly basic amino acids (or polybasic site) around the SARS-CoV-2 spike protein cleavage site also suggests this.
The viral spike protein is cleaved into two parts - S1 and S2, which is required for efficient viral infection of host cells. In influenza viruses, the hemeagglutinin (HA) protein -- which is analogous to spike protein in coronaviruses -- also acquires the polybasic site as it evolves from low to highly pathogenic influenza viruses.
How do we stay prepared?
While only 14 per cent of known human pathogens are viruses, these constitute 44 per cent of emerging pathogens, pointing to the growing threat of viral diseases. Of the viruses that have emerged in over four decades, about 75 per cent have a zoonotic origin. To continue to understand current and future pathogenic viruses and to maintain a level of preparedness, we must routinely sample animal species and human environments. Any sudden increases in human disease and fevers of unknown origin are also good focal points. Respiratory viruses would be best surveyed via ongoing surveillance of influenza-like illnesses and pneumonia in humans.
Infectious disease mortality has decreased largely due to better medical care, drugs to combat infections and vaccines to prevent disease. However, outbreaks of infectious disease worldwide have increased steadily since the 1980s. A complex mixture of biological, environmental, socio-economic and political factors threaten to stall past gains in infectious disease mortality and morbidity. These include pathogen evolution, antimicrobial resistance, climate change, global warming and deforestation, vaccine hesitancy, increasing pollution and population density and geopolitical conflicts, to name just a few. At the same time, new tools being developed in life sciences, nanotechnology, communications, space technologies and other fields offer science-based solutions to better detect, prevent, cure and control infectious diseases.
What is very clear is that the COVID-19 pandemic marks a wake-up call to better manage our ecology, continue to develop better mitigation tools and to deploy them effectively. Any future disease threats will be far better managed with science and transparency, instead of politics and scare-mongering.
(The writer is CEO of the DBT/Wellcome Trust India Alliance. He was head of the Virology Group at ICGEB, New Delhi.)
A product of evolution
Let’s pause here for a moment to take account of the fact that this finding is strong evidence that SARS-CoV-2 is not the product of deliberate manipulation but evolution. Creation in a lab is abrupt not gradual. Further, logic demands that someone trying to engineer a coronavirus to cause disease in humans would have used the backbone of a virus known to cause human disease. They would not have used a bat or pangolin virus that was never associated with human disease.
It is improbable that SARS-CoV-2 emerged through laboratory manipulation of a related SARS-CoV-like coronavirus. Genetic data conclusively show that SARS-CoV-2 is not derived from any previously used virus backbone.
SARS-CoV-2: How could it have evolved?
Scientists have proposed a few scenarios to explain the origin of SARS-CoV-2. One suggests that it arose from natural selection in an animal host before zoonotic transfer, i.e. transfer from an animal host to a human. The other favours zoonotic transfer followed by natural selection in humans.
As many early cases of COVID-19 were linked to the wet animal market of Wuhan, the potential source may have been there. (A “wet market” is a place selling fresh meat, fish, produce, and other perishable goods as distinguished from “dry markets' that sell durable goods such as fabric and electronics). Given the high similarity to bat SARS-CoV-like coronaviruses, the progenitor (or parent) of the virus may have jumped from bats, and then evolved into a receptor-binding domain (RBD) that efficiently binds to human ACE2.
Alternatively, the virus could have jumped from pangolins, which have an RBD that is highly similar to SARS-CoV-2. In either case, the progenitor of the virus would have to evolve the polybasic cleavage site. This improves a virus’s ability to penetrate host cells, reproduce, and spread from host to host. No progenitor virus has been found in any animal that has both these characteristics – efficient RBD and polybasic cleavage site.
A progenitor may have jumped into humans and evolved an optimized RBD and polybasic cleavage site while transmitting undetected in humans. Once acquired, these adaptations would enable efficient human-to-human transmission allowing enough cases to accumulate and be visible to the public health surveillance system.
Over 25,000 SARS-CoV-2 genomes have been sequenced so far and all have these two genomic features, suggesting a common ancestor, which has not been found. Phylogenetic analyses time this most recent common ancestor to late November or early December 2019. Sequencing of banked respiratory disease samples from this period may provide some insights.
Could SARS-CoV-2 have evolved in a laboratory and escaped from it? After the emergence of SARS in 2003 and its link to bats, many laboratories around the world isolated and cultured bat coronaviruses to study their properties. Could repeated passage of bat SARS-CoV-like coronaviruses in cell culture have given rise to SARS-CoV-2? Theoretically, it is possible for viruses cultured repeatedly on human cells to have evolved a RBD that bound more efficiently to human ACE2. But again, no progenitor viruses have been described from such laboratory studies going back more than a decade.
The most likely scenario supported by currently available scientific evidence is that a progenitor virus jumped from bats to pangolins to humans and evolved the polybasic cleavage site during its human-to-human transmission. However, it should be pointed out that more scientific data could shift the balance in favour of another hypothesis.
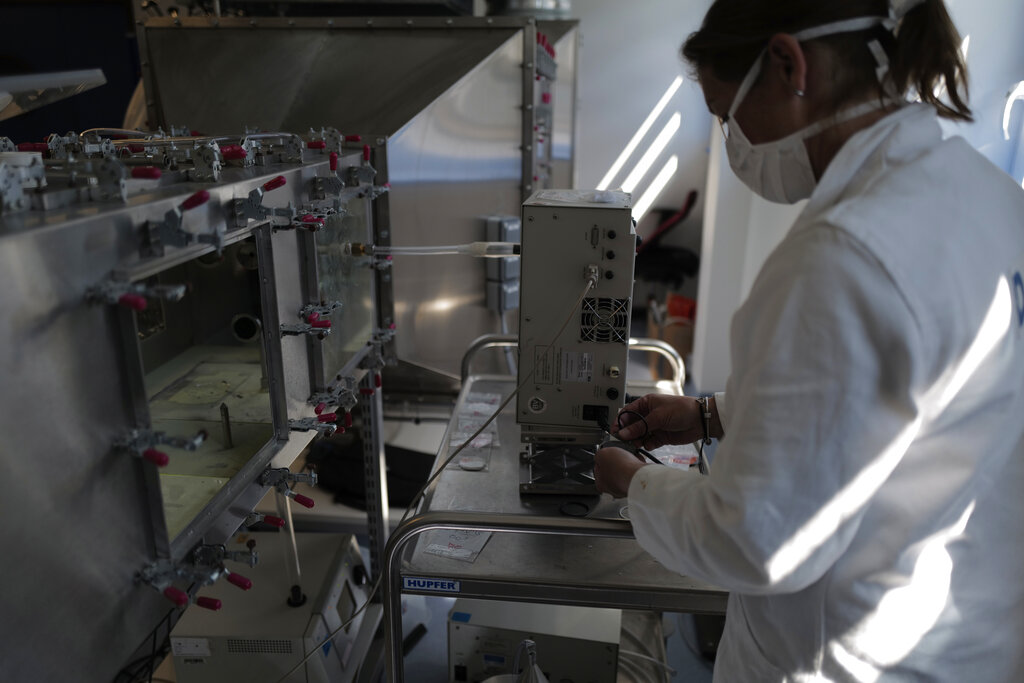

Technician of the French General Armament Directorate, DGA, specialized in research of Chemical Biological Radiological and Nuclear military protection gear analyzes public masks in their lab at Vert Le Petit, south of Paris, Wednesday, May 6, 2020 as a nationwide confinement continues to counter the Covid-19. (AP Photo/Francois Mori)
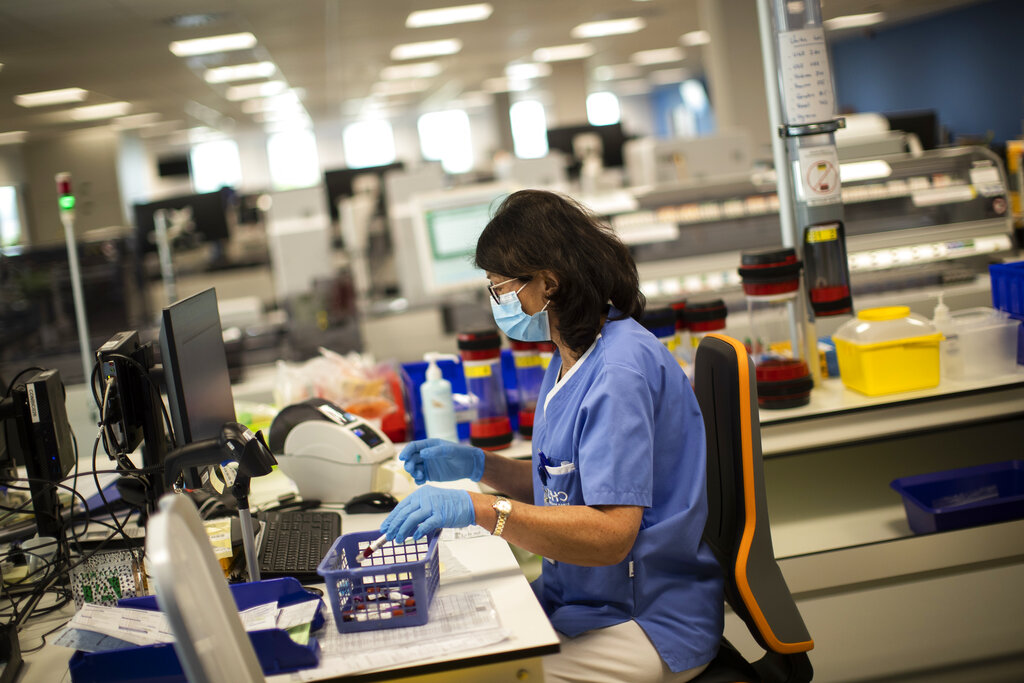

A member of medical personnel works with samples in a lab at the MontLegia CHC hospital, during a gradual lifting of a lockdown to prevent the spread of the coronavirus, in Liege, Belgium, Friday, May 8, 2020. (AP Photo/Francisco Seco)
An employee works in a research and development lab of Beijing Applied Biological Technologies, a firm which is developing Covid-19 molecular diagnostic test kits, during a government organized tour for journalists in Beijing, Thursday, May 14, 2020. (AP Photo/Mark Schiefelbein)